Volume-wise T2*/S0 estimation with t2smap#
Use tedana.workflows.t2smap_workflow() [DuPre et al., 2021] to calculate volume-wise T2*/S0,
as in Power et al. [2018] and Heunis et al. [2021].
import json
import os
from glob import glob
import matplotlib.pyplot as plt
import nibabel as nb
import numpy as np
from IPython import display
from book_utils import glue_figure, load_pafin
from nilearn import image, masking, plotting
from tedana import workflows
data_path = os.path.abspath('../data')
/mnt/c/Users/tsalo/Documents/ME-ICA/multi-echo-data-analysis/meda/lib/python3.10/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
data = load_pafin(data_path)
out_dir = os.path.join(data_path, "fit")
workflows.t2smap_workflow(
data['echo_files'],
data['echo_times'],
out_dir=out_dir,
mask=data['mask'],
prefix="sub-24053_ses-1_task-rat_dir-PA_run-01_",
fittype="loglin",
fitmode="ts",
overwrite=True,
)
out_files = sorted(glob(os.path.join(out_dir, "*")))
out_files = [os.path.basename(f) for f in out_files]
print("\n".join(out_files))
figures
sub-24053_ses-1_task-rat_dir-PA_run-01_S0map.nii.gz
sub-24053_ses-1_task-rat_dir-PA_run-01_T2starmap.nii.gz
sub-24053_ses-1_task-rat_dir-PA_run-01_dataset_description.json
sub-24053_ses-1_task-rat_dir-PA_run-01_desc-adaptiveGoodSignal_mask.nii.gz
sub-24053_ses-1_task-rat_dir-PA_run-01_desc-confounds_timeseries.tsv
sub-24053_ses-1_task-rat_dir-PA_run-01_desc-limited_S0map.nii.gz
sub-24053_ses-1_task-rat_dir-PA_run-01_desc-limited_T2starmap.nii.gz
sub-24053_ses-1_task-rat_dir-PA_run-01_desc-optcom_bold.nii.gz
sub-24053_ses-1_task-rat_dir-PA_run-01_desc-rmse_statmap.nii.gz
sub-24053_ses-1_task-rat_dir-PA_run-01_desc-tedana_registry.json
fig, axes = plt.subplots(figsize=(16, 16), nrows=3)
plotting.plot_carpet(
os.path.join(out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_desc-optcom_bold.nii.gz"),
axes=axes[0],
figure=fig,
)
axes[0].set_title("Optimally Combined Data", fontsize=20)
plotting.plot_carpet(
os.path.join(out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_T2starmap.nii.gz"),
axes=axes[1],
figure=fig,
)
axes[1].set_title("T2* Estimates", fontsize=20)
plotting.plot_carpet(
os.path.join(out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_S0map.nii.gz"),
axes=axes[2],
figure=fig,
)
axes[2].set_title("S0 Estimates", fontsize=20)
axes[0].xaxis.set_visible(False)
axes[1].xaxis.set_visible(False)
axes[0].spines["bottom"].set_visible(False)
axes[1].spines["bottom"].set_visible(False)
fig.tight_layout()
glue_figure("figure_volumewise_t2ss0_carpets", fig, display=False)
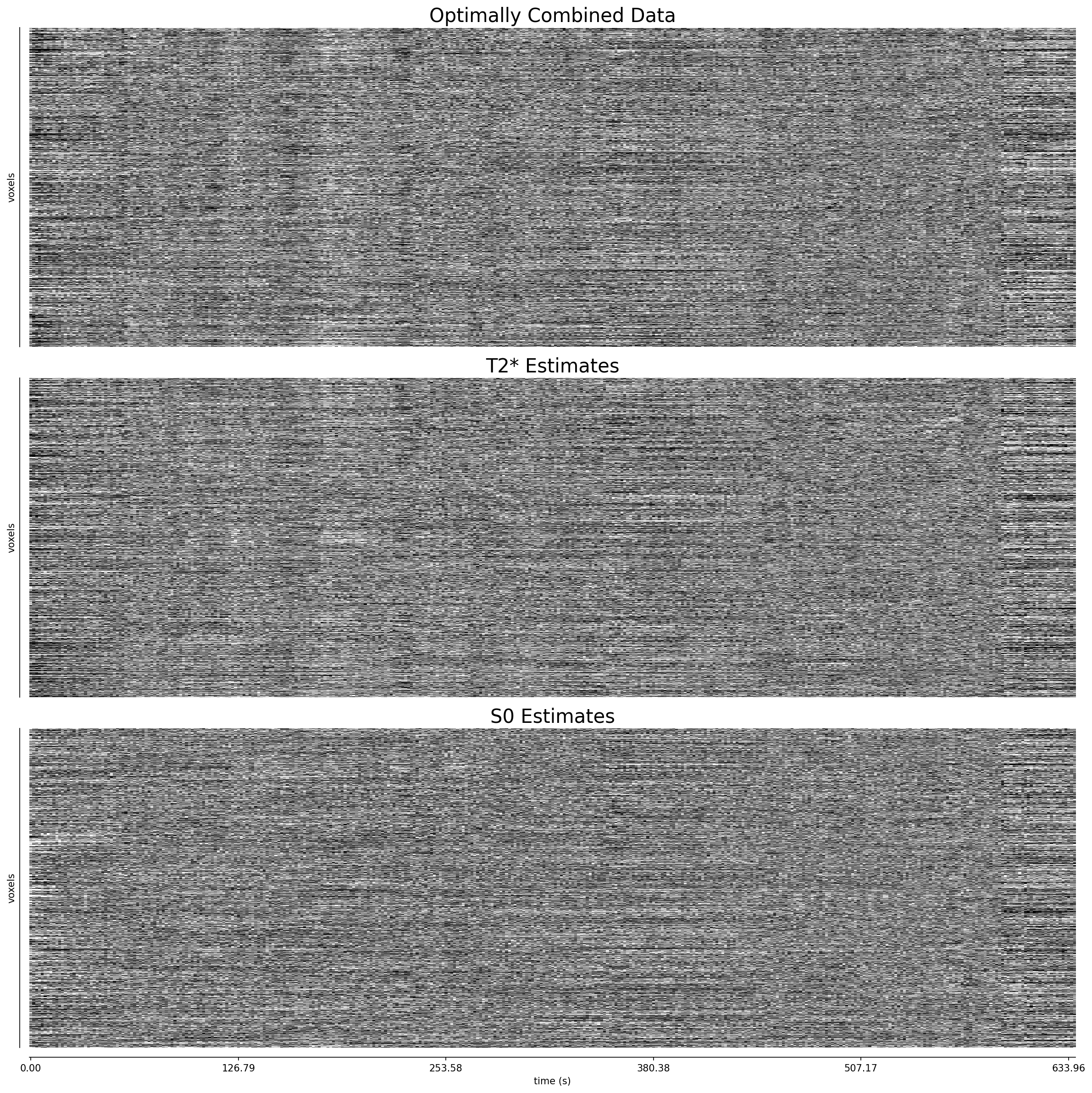

Fig. 44 Carpet plots of optimally combined data, along with volume-wise T2* and S0 estimates.#
fig, ax = plt.subplots(figsize=(16, 8))
in_file = image.mean_img(
os.path.join(out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_T2starmap.nii.gz")
)
arr = masking.apply_mask(in_file, data['mask'])
thresh = np.nanpercentile(arr, 98)
plotting.plot_stat_map(
in_file,
draw_cross=False,
bg_img=None,
figure=fig,
axes=ax,
cmap="Reds",
vmin=0,
vmax=thresh,
)
glue_figure("figure_mean_volumewise_t2s", fig, display=False)
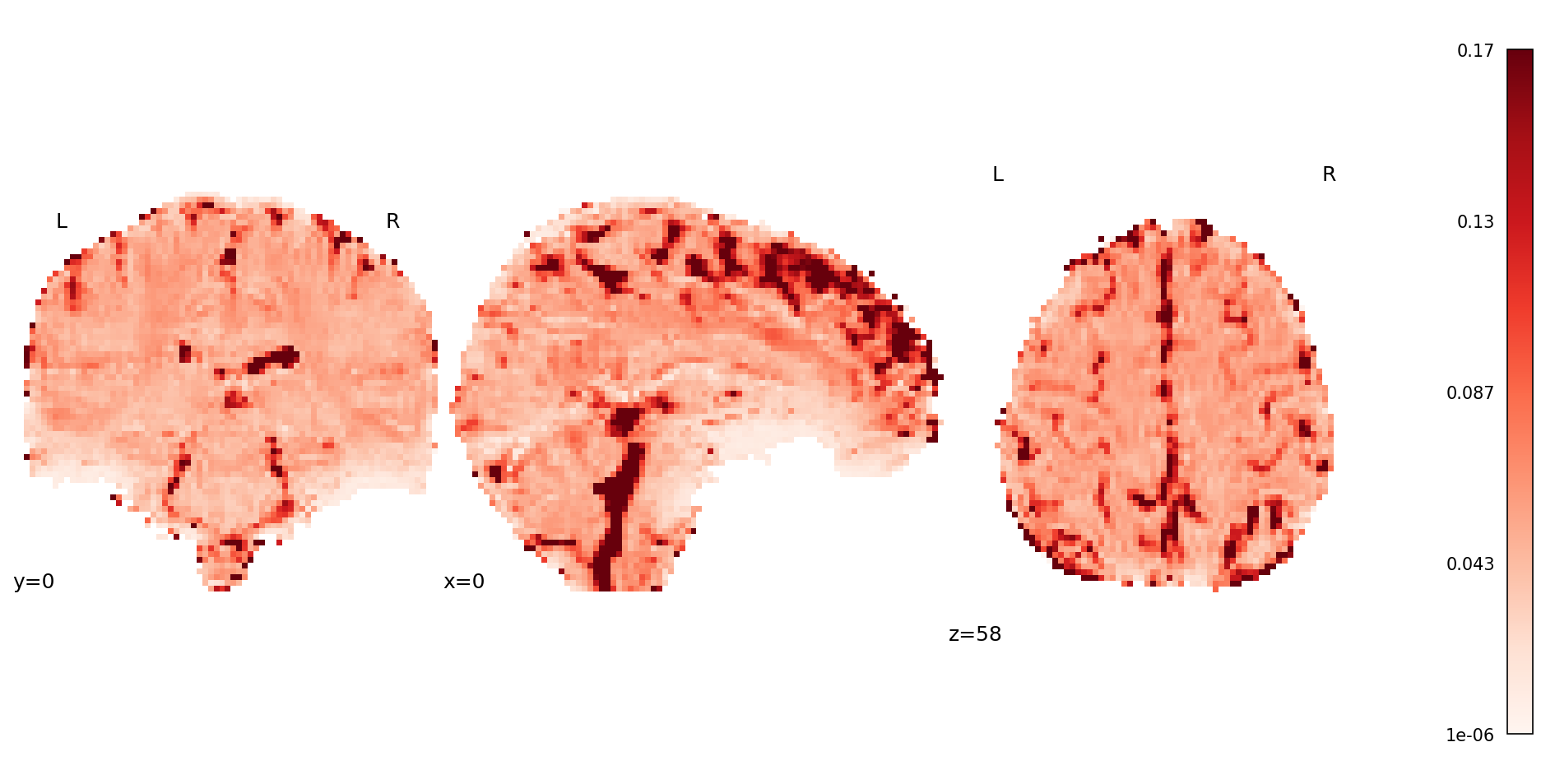
Fig. 45 Mean map from the volume-wise T2* estimation.#
fig, ax = plt.subplots(figsize=(16, 8))
in_file = image.mean_img(
os.path.join(out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_S0map.nii.gz")
)
arr = masking.apply_mask(in_file, data['mask'])
thresh = np.nanpercentile(arr, 98)
plotting.plot_stat_map(
in_file,
draw_cross=False,
bg_img=None,
figure=fig,
axes=ax,
cmap="Reds",
vmin=0,
vmax=thresh,
)
glue_figure("figure_mean_volumewise_s0", fig, display=False)
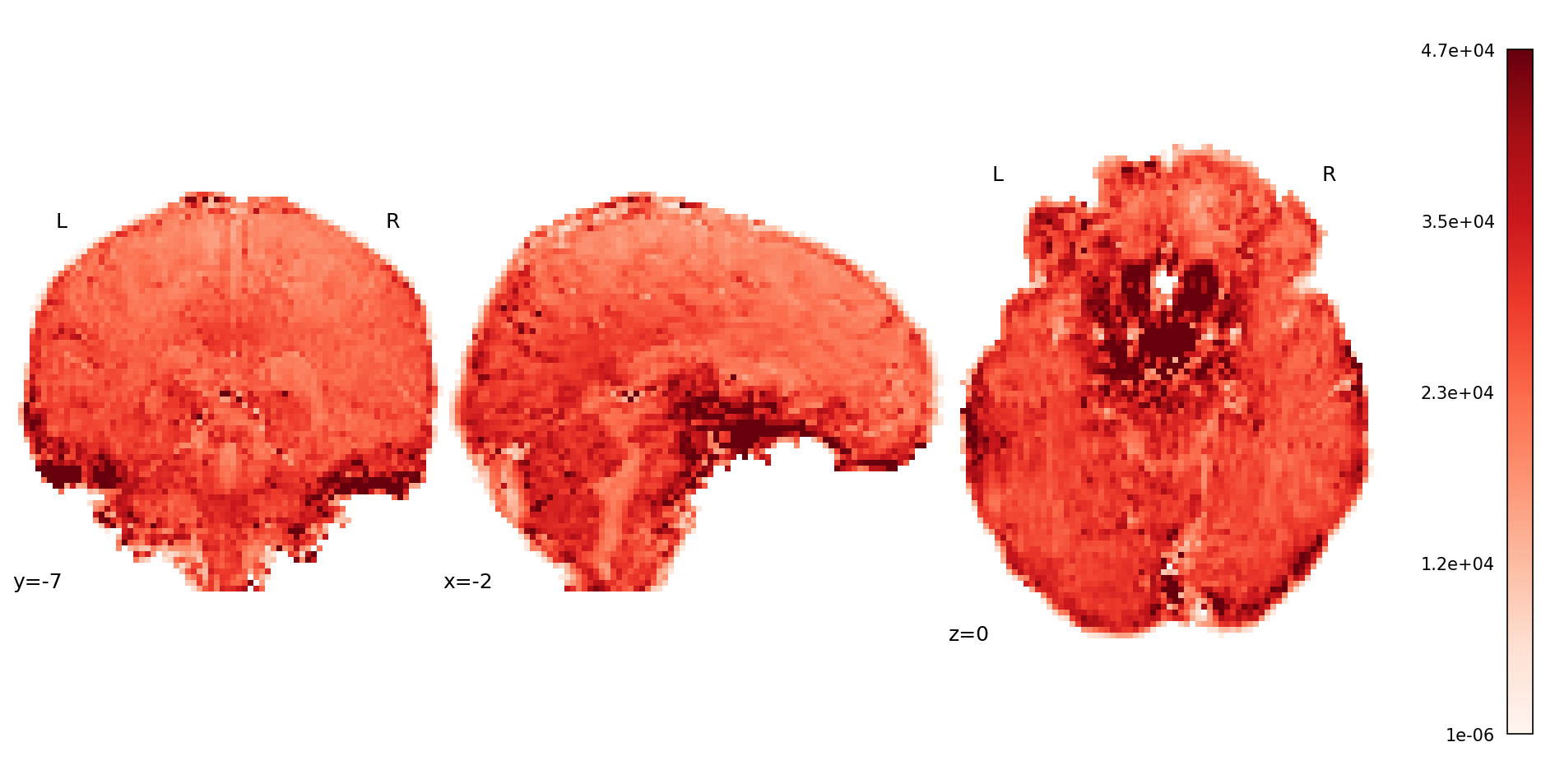
Fig. 46 Mean map from the volume-wise S0 estimation.#
fig, axes = plt.subplots(figsize=(16, 15), nrows=5)
plotting.plot_epi(
image.mean_img(data['echo_files'][0]),
draw_cross=False,
bg_img=None,
cut_coords=[-10, 0, 10, 20, 30, 40, 50, 60, 70],
display_mode="z",
axes=axes[0],
)
plotting.plot_epi(
image.mean_img(data['echo_files'][1]),
draw_cross=False,
bg_img=None,
cut_coords=[-10, 0, 10, 20, 30, 40, 50, 60, 70],
display_mode="z",
axes=axes[1],
)
plotting.plot_epi(
image.mean_img(data['echo_files'][2]),
draw_cross=False,
bg_img=None,
cut_coords=[-10, 0, 10, 20, 30, 40, 50, 60, 70],
display_mode="z",
axes=axes[2],
)
plotting.plot_epi(
image.mean_img(data['echo_files'][3]),
draw_cross=False,
bg_img=None,
cut_coords=[-10, 0, 10, 20, 30, 40, 50, 60, 70],
display_mode="z",
axes=axes[3],
)
plotting.plot_epi(
image.mean_img(
os.path.join(
out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_desc-optcom_bold.nii.gz"
)
),
draw_cross=False,
bg_img=None,
cut_coords=[-10, 0, 10, 20, 30, 40, 50, 60, 70],
display_mode="z",
axes=axes[4],
)
glue_figure("figure_mean_echos_and_optcom", fig, display=False)
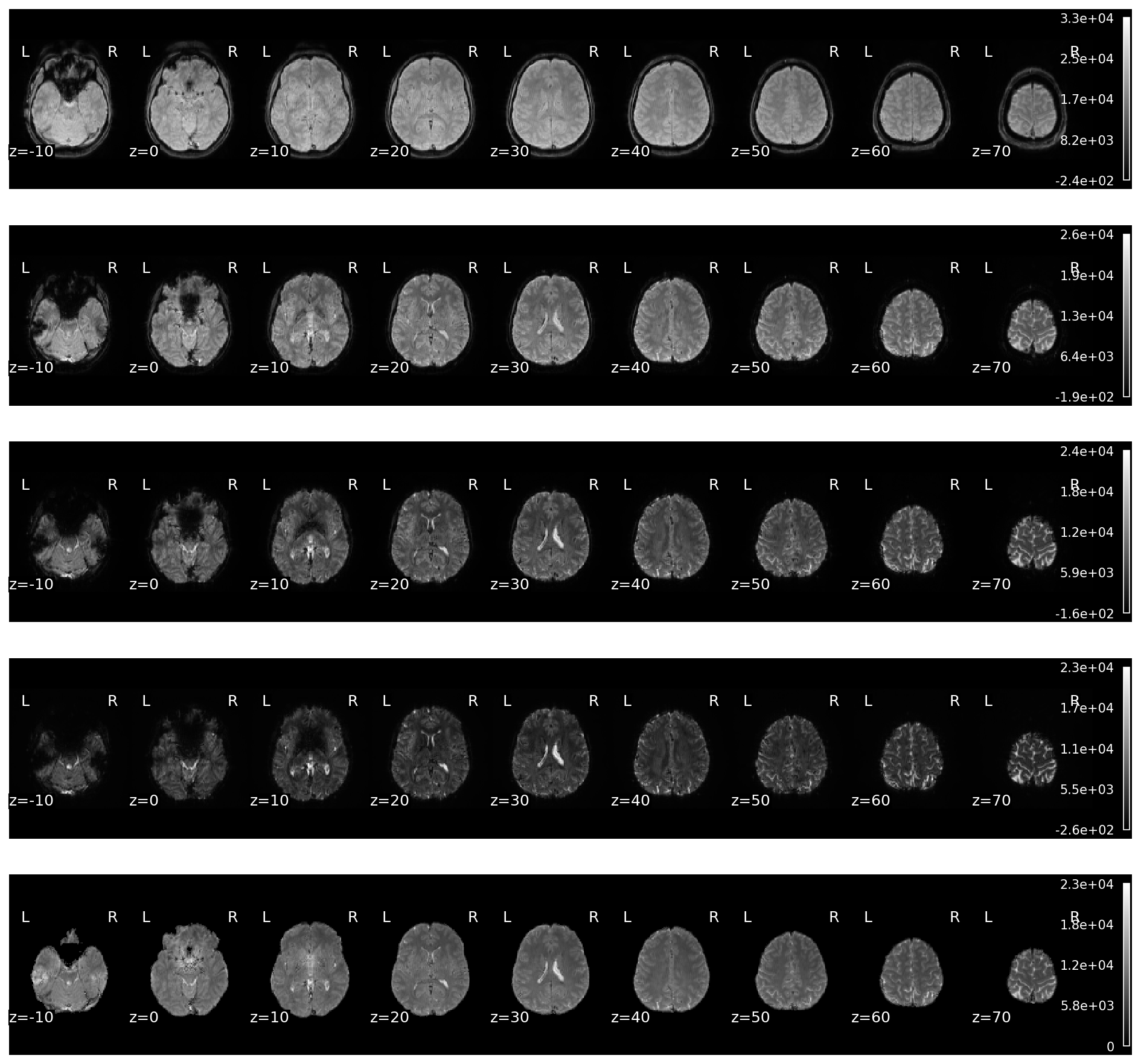
Fig. 47 Mean images of the echo-wise data and the optimally combined data.#
te30_tsnr = image.math_img(
"(np.nanmean(img, axis=3) / np.nanstd(img, axis=3)) * mask",
img=data['echo_files'][1],
mask=data['mask'],
)
oc_tsnr = image.math_img(
"(np.nanmean(img, axis=3) / np.nanstd(img, axis=3)) * mask",
img=os.path.join(
out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_desc-optcom_bold.nii.gz"
),
mask=data['mask'],
)
arr = masking.apply_mask(te30_tsnr, data['mask'])
thresh = np.nanpercentile(arr, 98)
arr = masking.apply_mask(oc_tsnr, data['mask'])
thresh = np.maximum(thresh, np.nanpercentile(arr, 98))
fig, axes = plt.subplots(figsize=(10, 8), nrows=2)
plotting.plot_stat_map(
te30_tsnr,
draw_cross=False,
bg_img=None,
threshold=0.1,
cut_coords=[0, 10, 10],
symmetric_cbar=False,
axes=axes[0],
cmap="Reds",
vmin=0,
vmax=thresh,
)
axes[0].set_title("TE30 TSNR", fontsize=16)
plotting.plot_stat_map(
oc_tsnr,
draw_cross=False,
bg_img=None,
threshold=0.1,
cut_coords=[0, 10, 10],
symmetric_cbar=False,
axes=axes[1],
cmap="Reds",
vmin=0,
vmax=thresh,
)
axes[1].set_title("Optimal Combination TSNR", fontsize=16)
glue_figure("figure_t2snr_te30_and_optcom", fig, display=False)
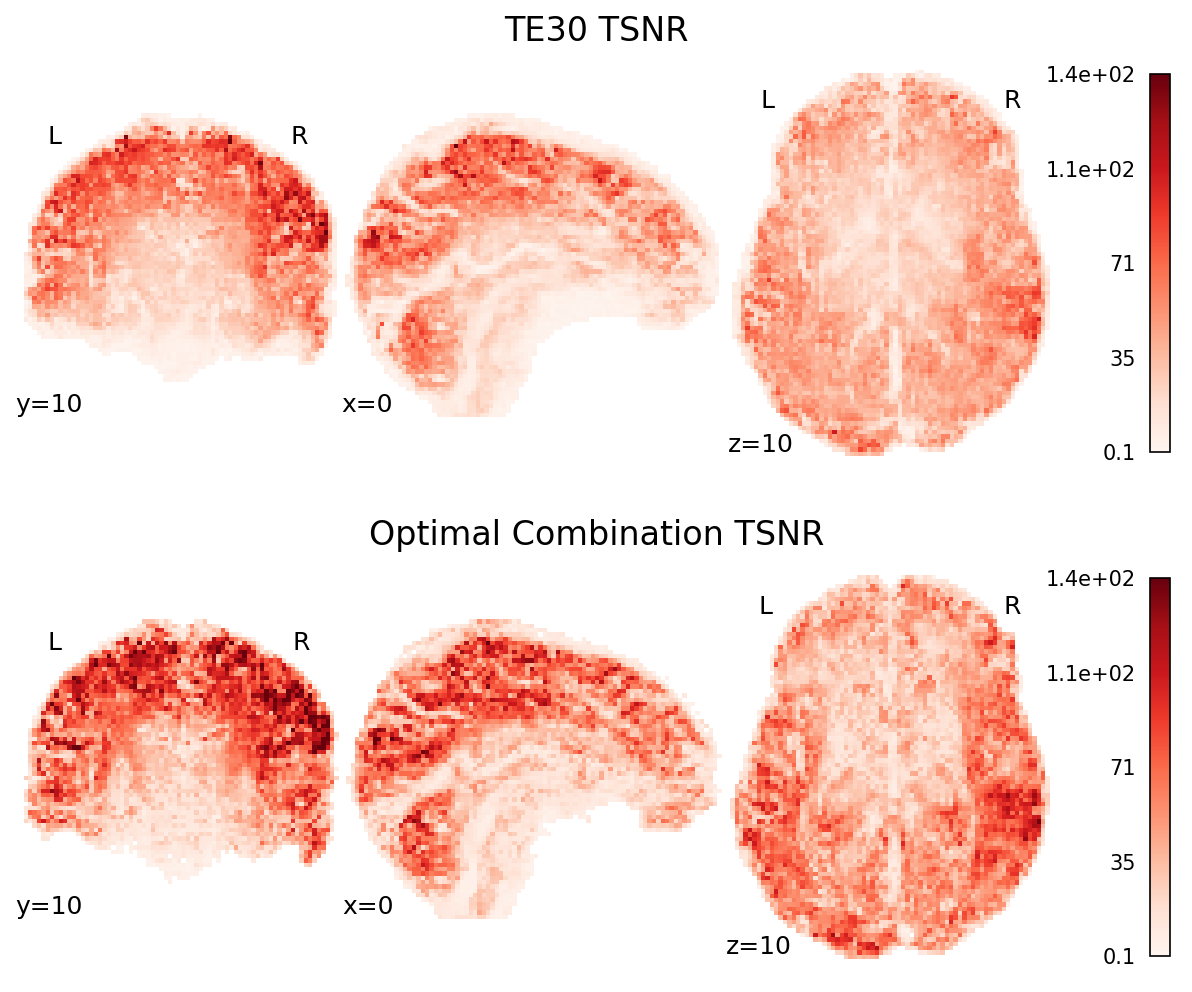
Fig. 48 TSNR map of the second echo (TE30) and the optimally combined data.#
fig, ax = plt.subplots(figsize=(16, 8))
plotting.plot_carpet(
data['echo_files'][1],
axes=ax,
)
glue_figure("figure_echo3_carpet", fig, display=False)
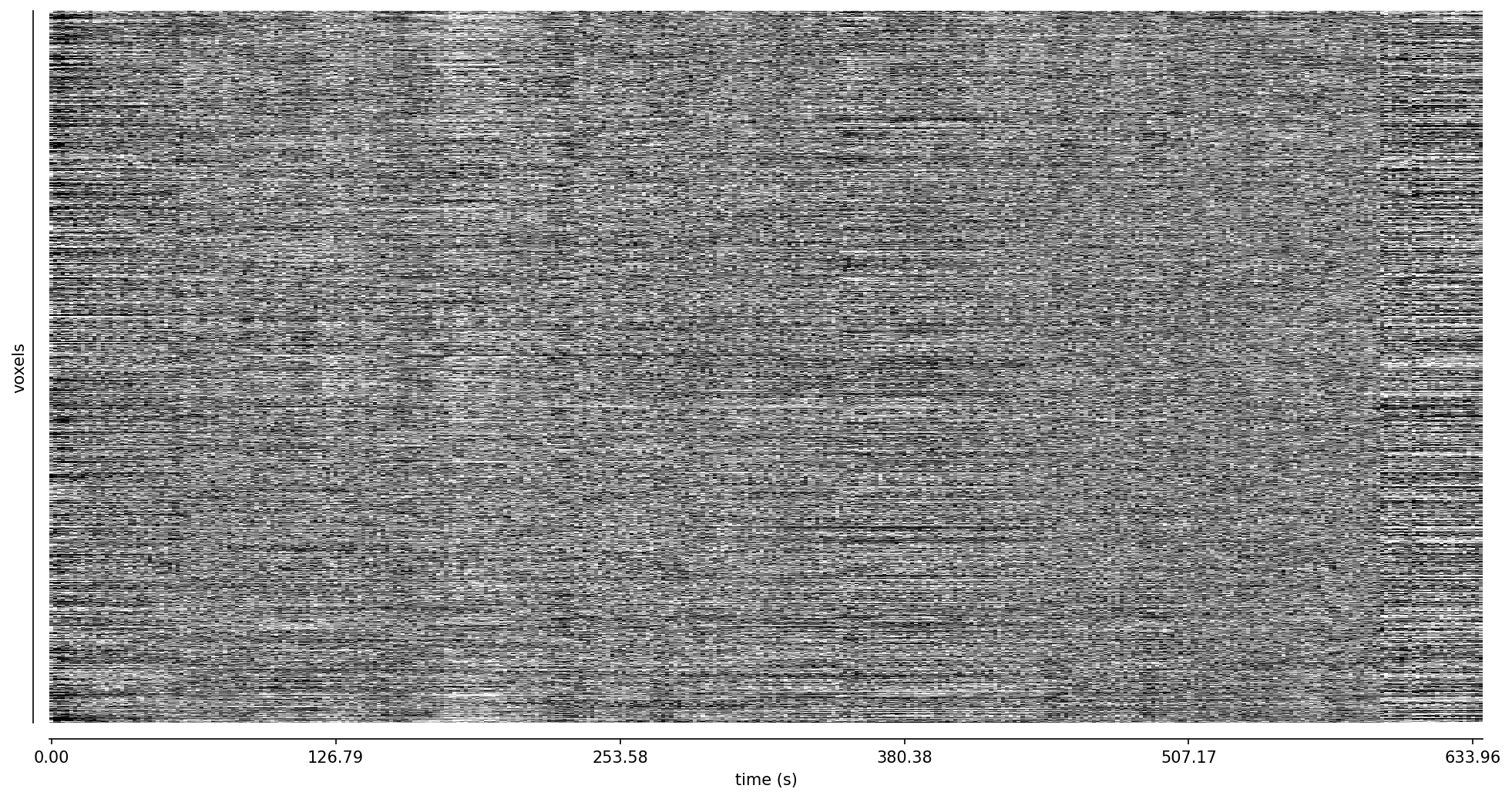
Fig. 49 Carpet plot of the third echo.#
fig, ax = plt.subplots(figsize=(16, 8))
plotting.plot_carpet(
os.path.join(out_dir, "sub-24053_ses-1_task-rat_dir-PA_run-01_desc-optcom_bold.nii.gz"),
axes=ax,
)
glue_figure("figure_carpet_optcom", fig, display=False)
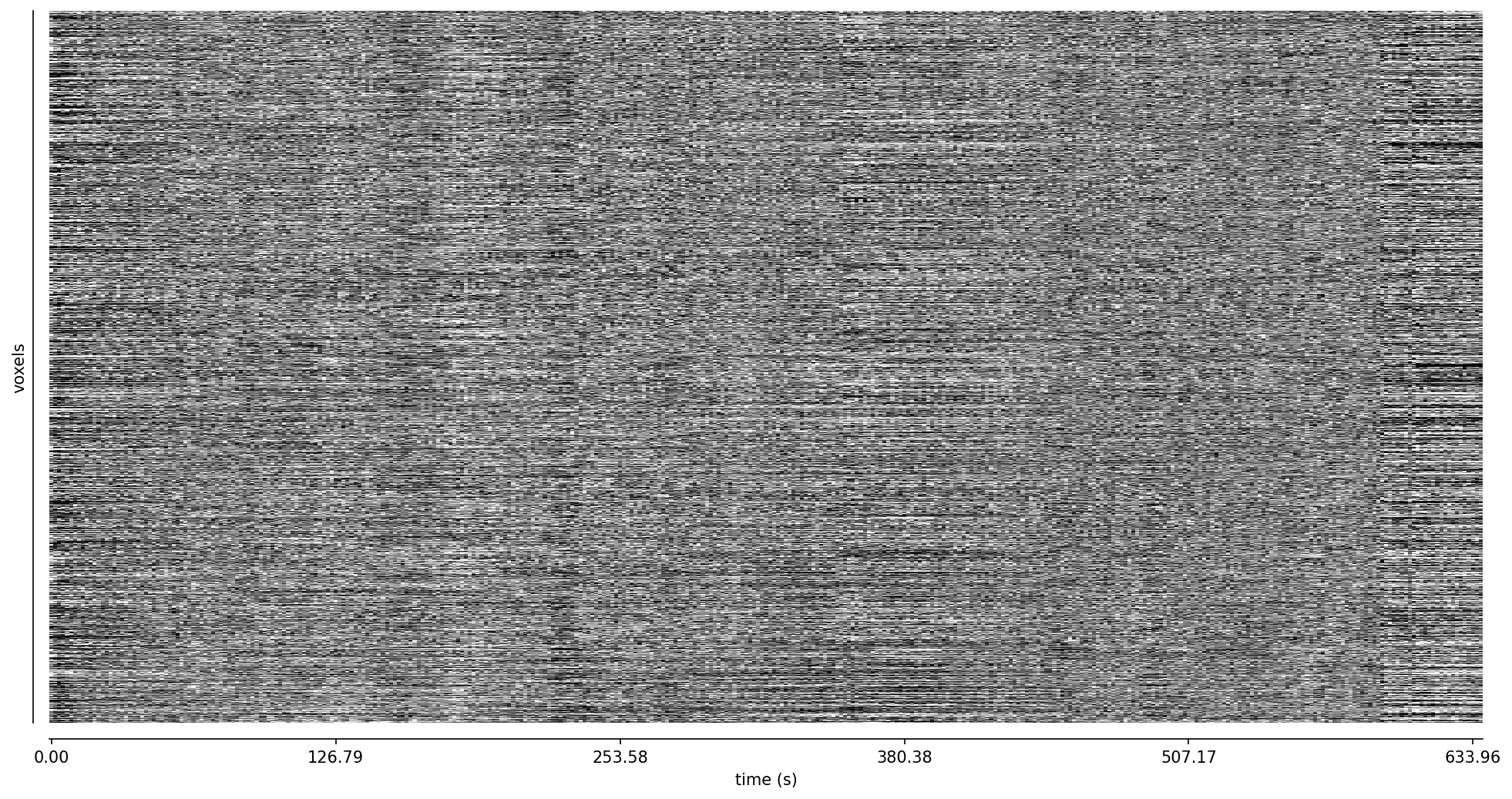
Fig. 50 Carpet plot of the optimally combined data.#

